About
I am a research scientist at Wageningen University & Research. My research interests include exploring -omics data to understand patterns and mechanisms of plant biotic interactions and co-evolution. Together with the WUR Bioinformatics group (Dr. Marnix Medema), Plant-Microbe interaction group (Prof. Saskia Van Wees/Prof. Corné Pieterse) at Utrecht university and Microbial ecology group at NIOO-KNAW (Prof. Jos Raaijmakers), I explore and implement innovative -omics integration strategies to map genes and their expression patterns to metabolites that play key roles in host-microbe interactions.
- Integrative -omics
- Bioinformatics
- Biotic interactions and co-evolution
PhD in Bioinformatics, 2014
University of Camerino Italy
MSc. in Bioinformatics, 2008
Nottingham Trent University UK
BSc in Biotechnology, 2006
University of Pune India
Research
In my current postdoctoral research at Wageningen University & Research and at Utrecht University, I investigate interactions between endophytes and their host-plants by integrating genomics, transcriptomics and metabolomics data. Please check the NWO Groot in the project section and a summary of my project in the form of a poster.
My overall research interest aims to integrate and explore -omics datasets (genomics, transcriptomics, epigenomics, proteomics and metabolomics) to understand patterns and mechanisms of biotic interactions and co-evolution.
My postdoctoral work at the University of Exeter and Rothamsted Research were mainly driven by key questions in resistance evolution of insect-pests to natural and synthetic insecticides. I used genome-wide investigations and computational approaches to answer such questions, ensuring these are relevant to a broad scientific community.
- Our work on brownplanthoppers in elucidating the role of the cytochrome P450 CYP6ER1 in resistance to neonicotinoid insecticides provided a novel example of gene duplication and neofunctionalization leading to resistance development in this species. Brown planthopper is the most devastating pest of south-east Asia and the findings of this study have helped in building suitable IPM strategies.
- Transcriptional plasticity of phytophagous generalist pests has always puzzled researchers. To further explore this, I have assembled the genome of a glasshouse whitefly, Trialeurodes vaporariorum, and helped perform transcriptomics in order to identify differences in expression of key genes on challenging host-plants. The findings in this work provide further insights in the ability of generalist pests to effectively reprogram genes expression during host-shift and the potential implication of transcriptional plasticity on their sensitivity to synthetic insecticides.
- My primary research has exploited cutting edge bioinformatics approaches to consistently progress beyond the state of the art. I have developed a novel software pipeline to electronically extend partial gene transcriptsup to full gene lengths, using raw RNA-seq data. This has helped characterize the correct number of genes, from different families, responsible for detoxifying allelochemicals in one of the solitary red mason bees.
- I also assembled the first genome sequence of a solitary bee Osmia bicornis and together with a PhD student showed that it lacks the CYP9Q subfamily of CYP450s but, despite this, it exhibits low acute toxicity to the neonicotinoid thiacloprid. Functional studies revealed that a P450 within the CYP9BU subfamily, with recent shared ancestry to the Apidae CYP9Q subfamily, metabolises thiacloprid in vitro and confers tolerancein vivo. This study is the landmark discovery of inherent tolerance of thiacloprid in solitary bees and can be leveraged to avoid negative pesticide impacts on these important pollinators.
- My most impactful academic output has been the work on aphid, Myzus persicae, and its host-shift to tobacco. In this work I used transcriptomics and genomics approaches to identify molecular signatures and genomic rearrangements that shape host mediated resistance evolution in aphids. My contribution in this work has laid the foundation of further experimental work that has provided key insights into the evolutionary processes facilitating insect adaptation to a novel host plant. Importantly, all genomic resources, along with four insect genomes and software developed by me, are used by many groups, extending beyond my field of expertise, to explore new and exciting avenues in molecular and evolutionary biology.
Experience
Featured Publications
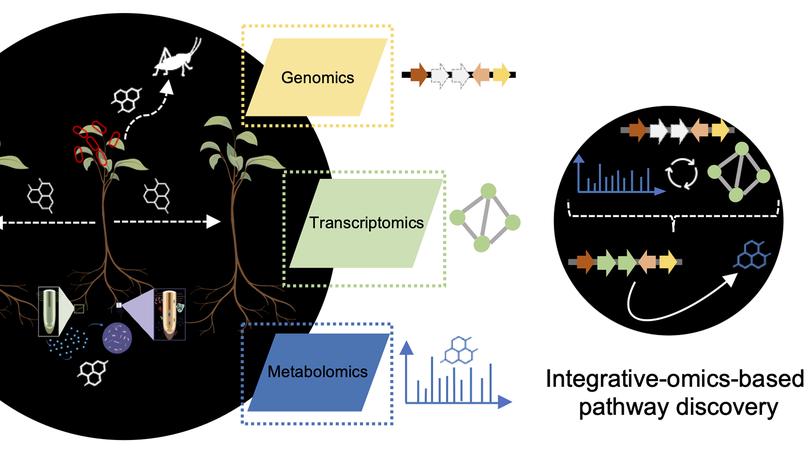
With the emergence of large amounts of omics data, computational approaches for the identification of plant natural product biosynthetic pathways and their genetic regulation have become increasingly important. While genomes provide clues regarding functional associations between genes based on gene clustering, metabolome mining provides a foundational technology to chart natural product structural diversity in plants, and transcriptomics has been successfully used to identify new members of their biosynthetic pathways based on coexpression. Thus far, most approaches utilizing transcriptomics and metabolomics have been targeted towards specific pathways and use one type of omics data at a time. Recent technological advances now provide new opportunities for integration of multiple omics types and untargeted pathway discovery. Here, we review advances in plant biosynthetic pathway discovery using genomics, transcriptomics, and metabolomics, as well as recent efforts towards omics integration. We highlight how transcriptomics and metabolomics provide complementary information to link genes to metabolites, by associating temporal and spatial gene expression levels with metabolite abundance levels across samples, and by matching mass-spectral features to enzyme families. Furthermore, we suggest that elucidation of gene regulatory networks using time-series data may prove useful for efforts to unwire the complexities of biosynthetic pathway components based on regulatory interactions and events.
Contact
- Droevendaalsesteeg 1, Wageningen, Wageningen University and Research 6708 PB
- Radix Building
- Thu-Fri 10.00 to 13:00
- Book an appointment
- DM Me